A growing list of biomarkers is beginning to drive targeted therapy in breast cancer, and clinicians can take advantage of these assays to make treatment selections, Debu Tripathy, MD, Professor of Medicine and Chair of Breast Medical Oncology at The University of Texas MD Anderson Cancer Center, told attendees at the 2019 Miami Breast Cancer Conference.1
As breast cancer tumors evolve over time, multiple “drivers” may be operational and may change dynamically. Functional genomic and expression signatures or consequences (such as mismatch repair) may identify patients who may respond to specific targeted agents. The serial assessment of these biomarkers can be clinically important, he said.
What Biomarkers Are Most Targetable?
In general, the types of lesions that are most targetable are those that lead to a gain of function. They include activating mutations (single nucleotide variants, fusions); amplifications (copy number variations); and phenotypic changes that reveal the behavior of a cell, such as DNA repair (mismatch-repair genes and increased mutational burden).

With next-generation sequencing, we provide a decision support framework. Patients receive a list of available drugs that might work in their situation, with links to phase I clinical trials.— Debu Tripathy, MD
Tweet this quote
“These are the genes we are most interested in finding,” Dr. Tripathy said. “Typically, they are going to be kinases that are activated, such as epidermal growth factor receptor (EGFR) in lung cancer, or they can be amplifications, such as HER2, where an excess amount of oncoprotein is made. Those are easier to target.”
On the other hand, mutations that cause loss of function, such as the tumor suppressor gene p53, are difficult to target. They can be deletions, frameshift and other loss-of-function mutations, and germline mutations (such as BRCA1 and BRCA2). Unfortunately, in breast cancer, targetable genomic lesions are few in number, he added.
Decision Support Framework for Investigational Therapies
Dr. Tripathy described the decision support framework used at MD Anderson Cancer Center for in-house sequencing and choice of treatment.2 The center uses a next-generation sequencing panel that covers about 150 genes and identifies mutations, copy number changes, and fusions. The functional consequences of these alterations are then determined, based on preclinical and clinical data.
If the functional alteration is in a driver gene, the framework identifies the relevant targeted agents and assesses the evidence for using a particular drug in the context of that patient. Where there is level I evidence, clinicians can select an approved therapy that is genomically matched. Where there is level II or III evidence, they can find a clinical trial using genotype-relevant drugs. The framework prioritizes mutations and targets them to identify the optimal treatment for a given patient, according to Dr. Tripathy.
“So, with next-generation sequencing, we provide a decision support framework. Patients receive a list of available drugs that might work in their situation, with links to phase I clinical trials as well,” he explained.
To date, MD Anderson has received 1,669 requests for annotation of 4,084 alterations across 49 tumor types for 1,197 patients. Requests are most common for colorectal, sarcoma, breast, and lung cancers. Of the 4,084 alterations (annotations), 78% (across all tumor types) were found to have an actionable gene, and about 20% of patients annotated were enrolled in a genotype-matched trial.
“For breast cancer, we’re actually able to match most of our patients to a phase I trial,” he reported. “What we haven’t shown yet is how our phase I trials are performing and how they performed before we were routinely doing next-generation sequencing. We are conducting that analysis now.”
Hints of Success for Novel Targets
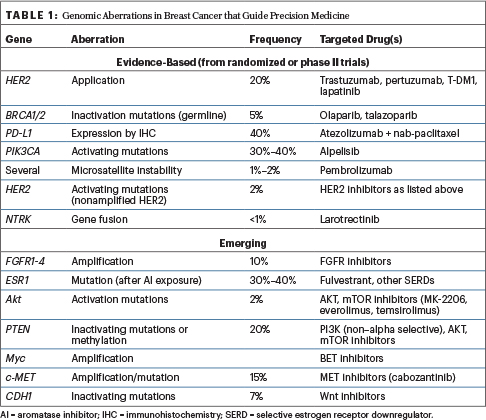
In retrospective analyses of all tumor types, genome-driven treatment has been shown to yield relatively high response rates, with a median response exceeding 2 years (the highest being with certain leukemias). Which alterations in breast cancer will be next? The list of genomic aberrations that actually guide precision medicine in breast cancer is constantly changing, but Dr. Tripathy shared the current and emerging landscape (Table 1).
PIK3CA is a common mutation in breast cancer, especially in hormone receptor–positive patients. Recent data from the SOLAR-1 trial confirmed that the alpha-selective PI3 kinase inhibitor alpelisib, when combined with fulvestrant, offers a clear progressive-free survival benefit compared with -placebo.3 This benefit was seen in PIK3CA mutation–positive cases alone, where the risk of disease progression was reduced by 52%; therefore, the drug will likely be approved in that subset alone.
Similarly, in patients with AKT E17–mutant triple-negative breast cancer, the AKT inhibitor ipatasertib was associated with a 56% reduction in the risk of disease progression, when added to paclitaxel compared with placebo, in a randomized phase II study.4,5 A phase III study is now underway.
“We have considerable experience at our institution with metaplastic breast cancers, which tend to have PI3 kinase pathway aberrations as well as activation of certain angiogenic tyrosine kinases,” he said. To address this scenario, Dr. Stacy Moulder and colleagues are using the combination of liposomal doxorubicin, bevacizumab, and everolimus; they are seeing response rates of around 30% in pathway-activated patients.6
“This is very different from what we have seen with chemotherapy,” he said. “We are now testing this in the neoadjuvant setting, to see whether we can actually induce responses and complete pathologic responses in patients who are relatively refractory to initial treatment with doxorubicin/cyclophosphamide.”
HER2 mutations (not amplifications but point mutations that activate the kinase) can be inhibited by kinase inhibitors, such as neratinib. In a recent update involving patients with HER2 mutations, responses to neratinib were seen in about 30% of patients.7 More recently, neratinib has been combined with fulvestrant in previously treated hormone receptor–positive patients with HER2 mutations, producing responses in the 40% range.8
Fibroblast growth factor receptor (FGFR) mutations (FGFR1–4) occur in 10% of breast cancers. An effective FGFR inhibitor is lacking for breast cancer, but several novel pan-FGFR inhibitors are now in development. “We are hoping we can improve the situation, because it is one of the more common mutations we see,” he commented.
Precision Basket Trials in the United States
Precision basket trials are determining which targeted treatments yield the best results in patients with alterations across tumor types. The “signal-finding” National Cancer Institute’s Molecular Analysis for Therapy Choice (NCI-MATCH) trial enrolled 6,000 mostly heavily pretreated patients with a variety of genomic aberrations; they were then assigned to about 30 different novel agents. For example, patients harboring AKT mutations have been treated with the AKT inhibitor capivasertib; response rates of 23% have been observed, and hormone receptor–positive patients have derived significant benefit, he said.
In the coming year, I suspect we will have our first genomically defined drug; soon thereafter, many more will follow.— Debu Tripathy, MD
Tweet this quote
The ongoing ASCO-sponsored TAPUR trial matches genomic lesions to drugs that have already been approved by the U.S. Food and Drug Administration (FDA). The trial has opened broadly across many community sites, as an effort to bring in practices that are not as active in early-phase trials or have in-house sequencing capacity, he said.
The MyPathway trial is evaluating treatment for patients with alterations in HER2 in cancers other than breast (trastuzumab plus pertuzumab) plus in BRAF (vemurafenib plus cobimetinib), smoothened (vismodegib), EGFR (erlotinib), anaplastic lymphoma kinase (ALK; alectinib), as well as copy number gain in programmed cell death protein ligand 1, mismatch repair deficiency (dMMR), microsatellite instability–high, and high tumor mutational burden (atezolizumab). Findings for HER2 overexpression and amplification in colorectal, bladder, and biliary cancers as well as for the BRAF V600E mutation in various tumor types were recently published.9
“By selecting these patients, responses are seen,” Dr. Tripathy noted. “It’s not always a home run, but responses are probably higher than if we randomly assigned patients to these targeted inhibitors.”
dMMR and Tumor Mutational Burden
The first tumor-agnostic FDA approval was for pembrolizumab in dMMR, on the basis of data first reported for colorectal cancer. The strong activity of checkpoint inhibitors in the dMMR subset led to a basket trial of patients with multiple tumor types expressing dMMR, with a 40% response rate shown.
“This was really astounding for a collection of tumors where response rates to pembrolizumab are typically much lower,” he said. Although dMMR/microsatellite instability–high disease is uncommon in breast cancer (< 2%), pembrolizumab can be used in these patients.
Tumor mutational burden is another overall phenotype being explored for targeted treatment, but there has been little to report in breast cancer. In lung cancer, tumor mutational burden does correlate positively with immunotherapy outcomes.
Applying Genomics in Practice
“In today’s practice, a few standard therapies ‘match’ specific targets, but it is more for the clinical trial candidate that sequencing and other ‘omic’ analyses can lead to new avenues,” Dr. Tripathy said in conclusion. “What we are doing in our practice is sequencing patients when they are at a point in time where they would typically be going on to clinical trials…. In the coming year, I suspect we will have our first genomically defined drug; soon thereafter, many more will follow.” ■
DISCLOSURE: Dr. Tripathy has served as a consultant for Nektar, Novartis, and Pfizer, and has received research support from Novartis.
REFERENCES
1. Tripathy D: Biomarker-directed therapy for metastatic breast cancer. 2019 Miami Breast Cancer Conference. Invited Lecture. Presented March 8, 2019.
2. Meric-Bernstam F, Johnson A, Holla V, et al: A decision support framework for genomically informed investigational cancer therapy. J Natl Cancer Inst 107:djv098, 2015.
3. André F, Ciruelos EM, Rubovszky G, et al: Alpelisib + fulvestrant for advanced breast cancer: Results of the phase 3 SOLAR-1 trial. ESMO 2018 Congress. Abstract LBA3. Presented October 20, 2018.
4. Hyman DM, Smyth LM, Donoghue MTA, et al: AKT inhibition in solid tumors with AKT1 mutations. J Clin Oncol 35:2251-2259, 2017.
5. Dent R, Im S-A, Espie M, et al: Overall survival update of the double-blind placebo-controlled randomized phase 2 LOTUS trial of first-line ipatasertib + paclitaxel for locally advanced/metastatic triple-negative breast cancer. 2018 ASCO Annual Meeting. Abstract 1008. Presented June 3, 2018.
6. Basho RK, Gilcrease M, Murthy RK, et al: Targeting the PI3K/AKT/mTOR pathway for the treatment of mesenchymal triple-negative breast cancer: Evidence from a phase 1 trial of mTOR inhibition in combination with liposomal doxorubicin and bevacizumab. JAMA Oncol 3:509-515, 2017.
7. Ma CX, Bose R, Gao F, et al: Neratinib efficacy and circulating tumor DNA detection of HER2 mutations in HER2 nonamplified metastatic breast cancer. Clin Cancer Res 23:5687-5695, 2017.
8. Smyth LM, Piha-Paul SA, Saura C, et al: Neratinib + fulvestrant for HER2-mutant, HR-positive, metastatic breast cancer: Updated results from the phase 2 SUMMIT trial. 2018 San Antonio Breast Cancer Symposium. Abstract PD3-06. Presented December 6, 2018.
9. Hainsworth JD, Meric-Bernstam F, Swanton C, et al: Targeted therapy for advanced solid tumors on the basis of molecular profiles: Results from MyPathway, an open-label, phase IIa multiple basket study. J Clin Oncol 36:536-542, 2018.

